Today we’re going to talk about another nonparametric test: the Moses Test for equal variability!
When Would You Use It?
The Moses Test for equal variability is a nonparametric test used to determine if two independent samples represent two populations with different variances.
What Type of Data?
The Moses Test for equal variability requires ordinal data.
Test Assumptions
- Each sample is a simple random sample from the population it represents.
- The two samples are independent.
Test Process
Step 1: Formulate the null and alternative hypotheses. The null hypothesis claims that the two population variances are equal. The alternative hypothesis claims otherwise (one variance is greater than the other, or that they are simply not equal).
Step 2: Compute the test statistics: U1 and U2. Since this is best done with data, please see the example shown below to see how this is done. [Note that the test statistic calculations are exactly the same as for the Mann-Whitney U test. The only thing that differs is the ranking procedure.]
Step 3: Obtain the critical value. Unlike most of the tests we’ve done so far, you don’t get a precise p-value when computing the results here. Rather, you calculate your U values and then compare them to a specific value. This is done using a table (such as the one here). Find the number at the intersection of your sample sizes for both samples at the specified alpha-level. Compare this value with the smaller of your U1 and U2 values.
Step 4: Determine the conclusion. If your test statistic is equal to or less than the table value, reject the null hypothesis. If your test statistic is greater than the table value, fail to reject the null (that is, claim that the variances are equal in the population).
Example
Today’s data come from my 2012 music selection (I’ll use this data next week, too!). I wanted to see if the median play counts for two genres—pop and electronic—were the same. I chose these two because I think most of my favorite songs are of one of the two genres. To keep things relatively simple for the example, I sampled n = 8 electronic songs and n = 8 pop songs. Set α = 0.05.
H0: σ2pop = σ2electronic
Ha: σ2pop ≠ σ2electronic
Here are the raw data:

The following tables show several different columns of information. I will explain the columns below.

The “Subsample” column: to obtain the rankings for this test, first divide the n1 scores in sample 1 into m1 subsamples (m1 > 1), with each subsample comprised of k scores. Then divide the n2 scores of sample 2 into m2 subsamples (m2 >2), with each subsample comprised of k scores. To form the subsamples, employ sampling without replacement within each of the samples. Ideally, m1, m2, and k should be chosen such that (m1)(k) = n1 and (m2)(k) = n2 if at all possible. Here, m1 = m2 = 4, and k = 2. The “subsample” columns list the four subsamples of play counts for each genre.
The second column contains the average of the k values for a given subset. It’s just the average of the values in the “subsample” column.
The third column contains the differences between each subsample value (X) and the subsample’s mean.
The fourth column is just the third column’s values squared.
The fifth column contains the sum of the values in the fourth column.
The sixth column contains the rank of the value in the fifth column over both genres. The smallest values is ranked as 1 and the largest is ranked as 8 (in this case).
To compute U1 and U2, use the following equations:

So here,
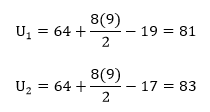
The test statistic itself is the smaller of the above values, so we get U = 81. In the table, the critical value for n1 = 8 and n2 = 8 and α = 0.05 for a two-tailed test is 13. Since U > 13, we fail to reject the null and retain the claim that the population variances are equal.
Example in R
x=read.table('clipboard', header=T) #data
attach(x)
elects=subset(x,genre=="Electronic")
pops=subset(x,genre=="Pop")
group1=matrix(rep(NaN,8),nrow=4)
group2=matrix(rep(NaN,8),nrow=4)
group1=as.matrix(sample(elects[,2],8,replace=F)) #subgroups for sample 1
sub1.1=group1[1:2,]
sub1.2=group1[3:4,]
sub1.3=group1[5:6,]
sub1.4=group1[7:8,]
group2=as.matrix(sample(pops[,2],8,replace=F)) #subgroups for sample 1
sub2.1=group2[1:2,]
sub2.2=group2[3:4,]
sub2.3=group2[5:6,]
sub2.4=group2[7:8,]
samps=rbind(sub1.1,sub1.2,sub1.3,sub1.4,sub2.1,sub2.2,sub2.3,sub2.4)
xbars=rep(NaN,8) #column 2
for(i in 1:8){
xbars[i]=mean(samps[i,])}
xbars=as.matrix(xbars)
diffs1=rep(NaN,8) #column 3
for(i in 1:8){
diffs1[i]=(samps[i,1]-xbars[i])}
diffs2=rep(NaN,8)
for(i in 1:8){
diffs2[i]=(samps[i,2]-xbars[i])}
diffs=cbind(diffs1,diffs2) #column 4
diffs2=diffs^2 #column 5
sumdif=rep(NaN,8)
for(i in 1:8){
sumdif[i]=diffs2[i,1]+diffs2[i,2]} #column 6
